Page 86 - MI-2-2
P. 86
Microbes & Immunity Big data and DNN-based DTI model in CHP
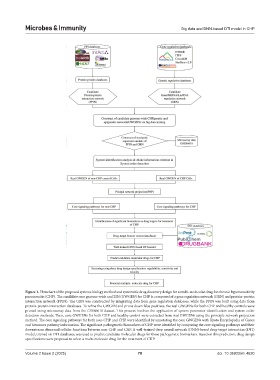
Figure 1. Flowchart of the proposed systems biology method and systematic drug discovery design for a multi-molecular drug for chronic hypersensitivity
pneumonitis (CHP). The candidate core genome-wide and EIN (GWGEN) for CHP is composed of a gene regulation network (GRN) and protein-protein
interaction network (PPIN). The GRN was constructed by integrating data from gene regulation databases, while the PPIN was built using data from
protein-protein interaction databases. To refine the GWGEN and prune down false positives, the real GWGENs for both CHP and healthy controls were
pruned using microarray data from the GSE86618 dataset. This process involves the application of system parameter identification and system order
detection methods. Then, core GWGENs for both CHP and healthy control were extracted from real GWGENs using the principle network projection
method. The core signaling pathways for both non-CHP and CHP were identified by annotating the core GWGENs with Kyoto Encyclopedia of Genes
and Genomes pathway information. The significant pathogenetic biomarkers of CHP were identified by comparing the core signaling pathways and their
downstream abnormal cellular functions between non-CHP and CHP. A well-trained deep neural network (DNN)-based drug-target interaction (DTI)
model, trained on DTI databases, was used to predict candidate molecular drugs for these pathogenetic biomarkers. Based on this prediction, drug design
specifications were proposed to select a multi-molecule drug for the treatment of CHP.
Volume 2 Issue 2 (2025) 78 doi: 10.36922/mi.4620