Page 57 - GPD-4-3
P. 57
Gene & Protein in Disease Insights from In situ spatial profiling
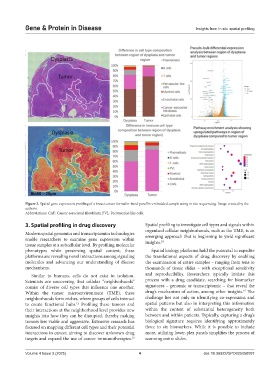
Figure 2. Spatial gene expression profiling of a breast cancer formalin-fixed paraffin-embedded sample using in situ sequencing. Image created by the
authors.
Abbreviations: CAF: Cancer-associated fibroblasts; PVL: Perivascular-like cells.
3. Spatial profiling in drug discovery Spatial profiling to investigate cell types and signals within
organized cellular neighborhoods, such as the TME, is an
Modern spatial genomics and transcriptomics technologies
enable researchers to examine gene expression within emerging approach that is beginning to yield significant
insights.
14
tissue samples at a subcellular level. By profiling molecular
phenotypes while preserving spatial context, these Spatial biology platforms hold the potential to expedite
platforms are revealing novel interactions among signaling the translational aspects of drug discovery by enabling
molecules and advancing our understanding of disease the examination of entire samples – ranging from tens to
mechanisms. thousands of tissue slides – with exceptional sensitivity
Similar to humans, cells do not exist in isolation. and reproducibility. Researchers typically initiate this
Scientists are uncovering that cellular “neighborhoods” process with a drug candidate, searching for biomarker
consist of diverse cell types that influence one another. signatures – genomic or transcriptomic – that reveal the
15
Within the tumor microenvironment (TME), these drug’s mechanism of action, among other insights. The
neighborhoods form niches, where groups of cells interact challenge lies not only in identifying co-expression and
to create functional hubs. Profiling these tumors and spatial patterns but also in interpreting this information
12
their interactions at the neighborhood level provides new within the context of substantial heterogeneity both
insights into how they can be disrupted, thereby making between and within patients. Typically, capturing a drug’s
tumors less viable and aggressive. Extensive research has biological signature requires identifying approximately
focused on mapping different cell types and their potential three to six biomarkers. While it is possible to include
interactions in cancer, aiming to discover unknown drug more, utilizing lower-plex panels simplifies the process of
targets and expand the use of cancer immunotherapies. scanning entire slides.
13
Volume 4 Issue 3 (2025) 4 doi: 10.36922/GPD025050007