Page 60 - GPD-4-3
P. 60
Gene & Protein in Disease Insights from In situ spatial profiling
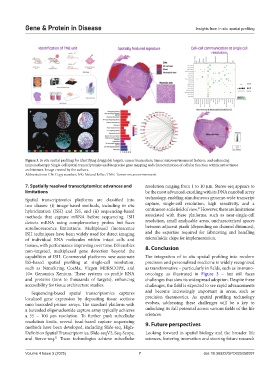
Figure 3. In situ spatial profiling for identifying druggable targets, cancer biomarkers, tumor microenvironment features, and enhancing
immunotherapy. Single-cell spatial transcriptomics enables precise gene mapping and characterization of cellular function within native tissue
architecture. Image created by the authors.
Abbreviations: CN: Copy number; NK: Natural killer; TME: Tumor microenvironment.
7. Spatially resolved transcriptomics: advances and resolution ranging from 1 to 10 μm. Stereo-seq appears to
limitations be the most advanced, excelling with its DNA nanoball array
Spatial transcriptomics platforms are classified into technology, enabling simultaneous genome-wide transcript
two classes: (i) image-based methods, including in situ capture, single-cell resolution, high sensitivity, and a
32
hybridization (ISH) and ISS, and (ii) sequencing-based centimeter-scale field of view. However, there are limitations
methods that capture mRNA before sequencing. ISH associated with these platforms, such as near-single-cell
detects mRNA using complementary probes but faces resolution, small analyzable areas, uncharacterized spaces
autofluorescence limitations. Multiplexed fluorescence between adjacent pixels (depending on channel distances),
ISH techniques have been widely used for direct imaging and the expertise required for fabricating and handling
of individual RNA molecules within intact cells and microfluidic chips for implementation.
tissues, with performance improving over time. ISS enables 8. Conclusion
non-targeted, multiplexed gene detection beyond the
capabilities of ISH. Commercial platforms now automate The integration of in situ spatial profiling into modern
ISS-based spatial profiling at single-cell resolution, precision and personalized medicine is widely recognized
such as NanoString CosMx, Vizgen MERSCOPE, and as transformative – particularly in fields, such as immuno-
10× Genomics Xenium. These systems co-profile RNA oncology, as illustrated in Figure 3 – but still faces
and proteins (tens to thousands of targets), enhancing challenges that slow its widespread adoption. Despite these
accessibility for tissue architecture studies. challenges, the field is expected to see rapid advancements
Sequencing-based spatial transcriptomics captures and become increasingly important in areas, such as
localized gene expression by depositing tissue sections precision theranostics. As spatial profiling technology
onto barcoded primer arrays. The standard platform with evolves, addressing these challenges will be a key to
a barcoded oligonucleotide capture array typically achieves unlocking its full potential across various fields of the life
a 55 – 100 μm resolution. To further push subcellular sciences.
resolution limits, several bead-based capture sequencing 9. Future perspectives
methods have been developed, including Slide-seq, High-
Definition Spatial Transcriptomics, Slide-seqV2, Seq-Scope, Looking forward in spatial biology and the broader life
and Stereo-seq. These technologies achieve subcellular sciences, fostering innovation and steering future research
31
Volume 4 Issue 3 (2025) 7 doi: 10.36922/GPD025050007