Page 63 - GPD-3-4
P. 63
Gene & Protein in Disease Drugs and immune infiltration in IPF
Table 1. Basic data from the three microarray databases derived from the GEO database
Dataset ID Platform Normal IPF Region
sample sample
GSE2052 GPL1739 Amersham Biosciences CodeLink Uniset Human I Bioarray 11 13 USA
GSE110147 GPL6244 [HuGene-1_0-st] Affymetrix Human Gene 1.0 ST Array [transcript (gene) version] 11 22 Canada
GSE53845 GPL6480 Agilent-014850 Whole Human Genome Microarray 4×44K G4112F (Probe Name version) 8 40 USA
Abbreviations: IPF: Idiopathic pulmonary fibrosis; GEO: Gene expression omnibus.
A B
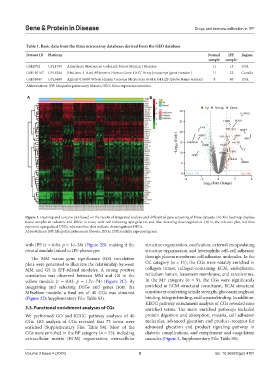
Figure 1. Heatmap and volcano plot based on the results of integrated analysis and differential gene screening of three datasets. (A) The heatmap displays
tissue samples in columns and DEGs in rows, with red indicating upregulation and blue denoting downregulation. (B) In the volcano plot, red dots
represent upregulated DEGs, whereas blue dots indicate downregulated DEGs.
Abbreviations: IPF: Idiopathic pulmonary fibrosis; DEGs: Differentially expressed genes.
with IPF (r = 0.84, p = 1e−28) (Figure 2B), making it the structure organization, ossification, external encapsulating
pivotal module linked to IPF phenotype. structure organization, and heterophilic cell–cell adhesion
The MM versus gene significance (GS) correlation through plasma membrane cell adhesion molecules. In the
plots were generated to illustrate the relationship between CC category (n = 11), the CGs were notably enriched in
MM and GS in IPF-related modules. A strong positive collagen trimer, collagen-containing ECM, endoplasmic
correlation was observed between MM and GS in the reticulum lumen, basement membrane, and sarcolemma.
yellow module (r = 0.93, p = 1.7e−74) (Figure 2C). By In the MF category (n = 9), the CGs were significantly
integrating and selecting DEGs and genes from the enriched in ECM structural constituent, ECM structural
MEyellow module, a final set of 40 CGs was obtained constituent conferring tensile strength, glycosaminoglycan
(Figure 2D; Supplementary File: Table S3). binding, integrin binding, and heparin binding. In addition,
KEGG pathway enrichment analysis of CGs revealed nine
3.3. Functional enrichment analyses of CGs enriched terms. The main enriched pathways included
We performed GO and KEGG pathway analyses of 40 protein digestion and absorption, malaria, cell adhesion
CGs. GO analysis of CGs revealed that 73 terms were molecules, advanced glycation end product–receptor for
enriched (Supplementary File: Table S4). Most of the advanced glycation end product signaling pathway in
CGs were enriched in the BP category (n = 53), including diabetic complications, and complement and coagulation
extracellular matrix (ECM) organization, extracellular cascades (Figure 3, Supplementary File: Table S5).
Volume 3 Issue 4 (2024) 5 doi: 10.36922/gpd.4101