Page 77 - BH-2-3
P. 77
Brain & Heart Lipids in ALS: Bad and beneficial
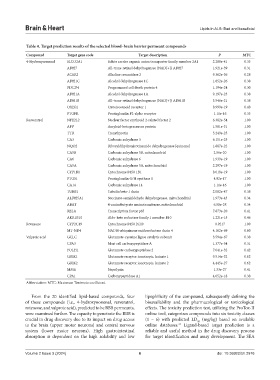
Table 4. Target prediction results of the selected blood–brain barrier permeant compounds
Compound Target gene code Target description P MTC
4-Hydroxynonenal SLCO2A1 Solute carrier organic anion transporter family member 2A1 2.288e-41 0.35
ADH7 All-trans-retinol dehydrogenase (NAD[+]) ADH7 1.921e-39 0.31
ACER2 Alkaline ceramidase 2 9.362e-36 0.28
ADH1C Alcohol dehydrogenase 1C 1.652e-26 0.38
PDCD4 Programmed cell death protein 4 1.194e-24 0.30
ADH1A Alcohol dehydrogenase 1A 9.197e-23 0.38
ADH1B All-trans-retinol dehydrogenase (NAD[+]) ADH1B 3.546e-21 0.38
OXER1 Oxoeicosanoid receptor 1 8.999e-19 0.48
PTGFR Prostaglandin F2-alpha receptor 1.11e-16 0.35
Resveratrol NFE2L2 Nuclear factor erythroid 2-related factor 2 6.382e-54 1.00
APP Amyloid-beta precursor protein 1.581e-31 1.00
TTR Transthyretin 5.249e-25 1.00
CA3 Carbonic anhydrase 3 6.151e-25 1.00
NQO2 Ribosyldihydronicotinamide dehydrogenase [quinone] 1.887e-22 1.00
CA5B Carbonic anhydrase 5B, mitochondrial 2.56e-20 1.00
CA6 Carbonic anhydrase 6 1.939e-19 1.00
CA5A Carbonic anhydrase 5A, mitochondrial 2.297e-19 1.00
CYP1B1 Cytochrome P450 1B1 8.018e-19 1.00
PTGS1 Prostaglandin G/H synthase 1 4.92e-17 1.00
CA14 Carbonic anhydrase 14 1.11e-16 1.00
TUBB1 Tubulin beta-1 chain 2.082e-47 0.38
ALDH5A1 Succinate-semialdehyde dehydrogenase, mitochondrial 1.973e-43 0.34
ABAT 4-aminobutyrate aminotransferase, mitochondrial 6.58e-28 0.34
RELA Transcription factor p65 7.873e-20 0.41
AKR1B10 Aldo-keto reductase family 1 member B10 1.221e-15 0.46
Rotenone CYP2C19 Cytochrome P450 2C19 0.9217 1.00
MT-ND4 NADH-ubiquinone oxidoreductase chain 4 4.162e-69 0.60
Valproic acid GCLC Glutamate-cysteine ligase catalytic subunit 3.594e-67 0.38
CPA3 Mast cell carboxypeptidase A 1.377e-54 0.31
FOLH1 Glutamate carboxypeptidase 2 7.041e-32 0.42
GRIK1 Glutamate receptor ionotropic, kainate 1 9.514e-32 0.62
GRIK2 Glutamate receptor ionotropic, kainate 2 4.445e-27 0.62
MME Neprilysin 1.33e-27 0.41
CPA1 Carboxypeptidase A1 4.052e-18 0.38
Abbreviation: MTC: Maximum Tanimoto coefficient.
From the 20 identified lipid-based compounds, four lipophilicity of the compound, subsequently defining the
of these compounds (i.e., 4-hydroxynonenal, resveratrol, bioavailability and the pharmacological or toxicological
rotenone, and valproic acid), predicted to be BBB permeants, effects. The toxicity prediction test, utilizing the ProTox-II
were examined further. The capacity to penetrate the BBB is online tool, categorizes compounds into six toxicity classes
crucial in drug discovery due to its impact on drug access (1 – 6) with predicted LD (mg/kg) based on available
50
to the brain (upper motor neurons) and central nervous online databases. Ligand-based target prediction is a
33
system (lower motor neurons). High gastrointestinal reliable and useful method in the drug discovery process
absorption is dependent on the high solubility and low for target identification and assay development. The SEA
Volume 2 Issue 3 (2024) 6 doi: 10.36922/bh.2976