Page 59 - GPD-2-2
P. 59
Gene & Protein in Disease Transfection methods for TK6 cells
A C
B D
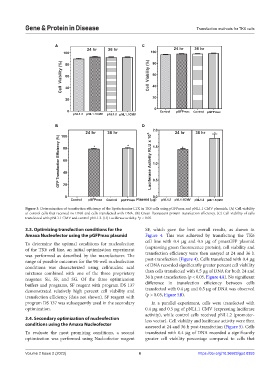
Figure 3. Determination of transfection efficiency of the lipofectamine LTX in TK6 cells using pGFPmax and pNL1.1 CMV plasmids. (A) Cell viability
of control cells that received no DNA and cells transfected with DNA. (B) Green fluorescent protein transfection efficiency. (C) Cell viability of cells
transfected with pNL1.1 CMV and control pNL1.2. (D) Luciferase activity. *p < 0.05.
3.3. Optimizing transfection conditions for the SF, which gave the best overall results, as shown in
Amaxa Nucleofector using the pGFPmax plasmid Figure 4. This was achieved by transfecting the TK6
To determine the optimal conditions for nucleofection cell line with 0.4 µg and 0.5 µg of pmaxGFP plasmid
of the TK6 cell line, an initial optimization experiment (expressing green fluorescence protein); cell viability and
was performed as described by the manufacturer. The transfection efficiency were then assayed at 24 and 36 h
range of possible outcomes for the 96-well nucleofection post-transfection (Figure 4). Cells transfected with 0.4 µg
conditions was characterized using cell/nucleic acid of DNA recorded significantly greater percent cell viability
mixtures combined with one of the three proprietary than cells transfected with 0.5 µg of DNA for both 24 and
reagents: SE, SF, and SG. Of the three optimization 36 h post-transfection (p < 0.05, Figure 4A). No significant
buffers and programs, SF reagent with program DS 137 difference in transfection efficiency between cells
demonstrated relatively high percent cell viability and transfected with 0.4 µg and 0.5 µg of DNA was observed
transfection efficiency (data not shown). SF reagent with (p > 0.05, Figure 5B).
program DS 137 was subsequently used in the secondary In a parallel experiment, cells were transfected with
optimization. 0.4 µg and 0.5 µg of pNL1.1 CMV (expressing luciferase
activity), while control cells received pNL1.2 (promoter-
3.4. Secondary optimization of nucleofection less vector). Cell viability and luciferase activity were then
conditions using the Amaxa Nucleofector assessed at 24 and 36 h post-transfection (Figure 5). Cells
To evaluate the most promising conditions, a second transfected with 0.4 µg of DNA recorded a significantly
optimization was performed using Nucleofector reagent greater cell viability percentage compared to cells that
Volume 2 Issue 2 (2023) 6 https://doi.org/10.36922/gpd.0353