Page 92 - GPD-3-3
P. 92
Gene & Protein in Disease Perineural invasion in prostate cancer
pathways activated only by common DEGs shared by both were upregulated only in perineural invasion-negative
Schwann cells and perineural invasion-negative samples, tumors (Figure 1A and B).
while regulation of actin cytoskeleton was activated only
by common DEGs shared by both Schwann cells and 3.5. Analysis of clock-related genes
perineural invasion-positive samples (Supplementary Since circadian entrainment is a pathway upregulated
File 1 Tables S5-S6). in perineural invasion-positive prostate tumors, we
associated DEGs to clock-related gene list. PER3, NR3C1,
3.3. Hallmarks of cancer analysis PPARGC1A, TIMP3, ID2, PDE6B, SLC25A10, and
In order to delineate the mechanism by which Schwann CAVIN1 were upregulated in perineural invasionnegative
cells aid in neoplastic development, we analyzed the tumors. We also observed upregulated genes in perineural
behavior of genes associated with cell differentiation invasion-positive tumors, such as PPARGC1A, TIMP3,
processes (GATA3, CDH1, and CDH2), apoptosis (CASP3, S100A8, ID2, DEFB1, AQP3, ASS1, PDE6B, NEFH, and
CASP9, BAX, and BCL2), motility (CXCR2, CXCL5, MMP9, CAVIN1 (Figure 1C and D). These DEG analysis outputs
and CCL12), and cell proliferation (MKI67). In perineural are supported by immunohistochemistry results (Figure 2).
invasion-negative tumors, GATA3, BCL2 and CXCR2
were upregulated, and GATA3 was upregulated, whereas 3.6. Methylation and copy number alteration
MMP9 was downregulated in perineural invasion-positive Methylation analysis of the GATA3, BCL2, CXCR2, MMP9,
samples (Figure 1A and B). NCAM1, NGFR, ROBO1, AQP3, NEFH, and PER3 genes from
tumor samples demonstrated that all of them were methylated
3.4. Analysis of genes involved in dedifferentiation in their promoter regions unlike those from normal tissues
of Schwann cells (data not shown). However, there was a positive correlation
The expression of cell differentiation maintenance proteins between methylation and MPZ and NR3C1 gene expression
(SOX10, S100, EGR2, MBP, ROBO1, ROBO2, SLIT2, (Figures 3 and 4, respectively). Copy number alteration
and MPZ) in Schwann cells is diminished after nerve data demonstrated that CDH1, CDH2, GFAP, PERP, and
damage, provoking cellular dedifferentiation. Immediately ROBO2 had a higher mRNA expression than normal tissues;
after injury, the cell body initiates a series of metabolic increased expression was associated with gain or amplification
responses, cytoplasmic reorganization and specific changes alterations in PRAD samples (Figure 5).
in gene expression (upregulation of SOX10, GAP43, S100,
NCAM1, NGFR1, and GFAP), collectively known as a 3.7. Correlation and prognostic analyses
neuronal reaction. 18,26,27 We observed increased expression In order to analyze the pathway by which Schwann
of NCAM1 and NGFR genes in both perineural invasion- cells induce neoplasm and their own cell proliferation
negative and -positive cancers, whereas ROBO1 and MPZ and migration, we evaluated the expression of AKT
A B
C
D
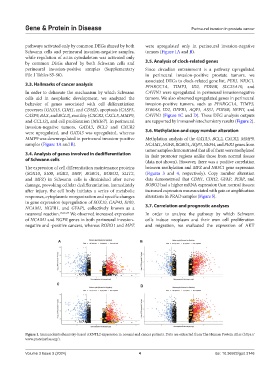
Figure 1. Immunohistochemistry-based ARNTL2 expression in normal and cancer patients. Data are extracted from The Human Protein Atlas (https://
www.proteinatlas.org/).
Volume 3 Issue 3 (2024) 4 doi: 10.36922/gpd.3146