Page 88 - GPD-3-4
P. 88
Gene & Protein in Disease A prediction on how Epimedii herba treat periodontitis
A B
C D
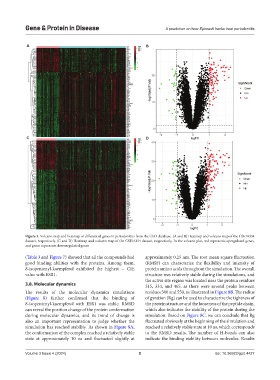
Figure 1. Volcano map and heatmap of differential genes in periodontitis from the GEO database. (A and B) Heatmap and volcano map of the GSE10334
dataset, respectively. (C and D) Heatmap and volcano map of the GSE16134 dataset, respectively. In the volcano plot, red represents upregulated genes,
and green represents downregulated genes
(Table 3 and Figure 7) showed that all the compounds had approximately 0.25 nm. The root mean square fluctuation
good binding abilities with the proteins. Among them, (RMSF) can characterize the flexibility and intensity of
8-isopentenyl-kaempferol exhibited the highest − CIE protein amino acids throughout the simulation. The overall
value with ESR1. structure was relatively stable during the simulations, and
the active site region was located near the protein residues
3.8. Molecular dynamics 315, 334, and 465, as there were several peaks between
The results of the molecular dynamics simulations residues 300 and 550, as illustrated in Figure 8B. The radius
(Figure 8) further confirmed that the binding of of gyration (Rg) can be used to characterize the tightness of
8-isopentenyl-kaempferol with ESR1 was stable. RMSD the protein structure and the looseness of the peptide chain,
can reveal the position change of the protein conformation which also indicates the stability of the protein during the
during molecular dynamics, and its trend of change is simulation. Based on Figure 8C, we can conclude that Rg
also an important representation to judge whether the fluctuated obviously at the beginning of the simulation and
simulation has reached stability. As shown in Figure 8A, reached a relatively stable state at 10 ns, which corresponds
the conformation of the complex reached a relatively stable to the RMSD results. The number of H-bonds can also
state at approximately 10 ns and fluctuated slightly at indicate the binding stability between molecules. Results
Volume 3 Issue 4 (2024) 5 doi: 10.36922/gpd.4427