Page 77 - GPD-4-3
P. 77
Gene & Protein in Disease Cullin family members as biomarkers in KIRC
A
B C
D E
F
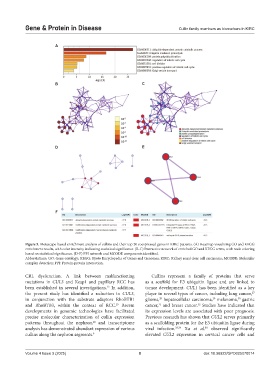
Figure 5. Metascape-based enrichment analysis of cullins and their top 20 coexpressed genes in KIRC patients. (A) Heatmap visualizing GO and KEGG
enrichment results, with color intensity indicating statistical significance. (B-C) Interactive network of enriched GO and KEGG terms, with node coloring
based on statistical significance. (D-F) PPI network and MCODE components identified.
Abbreviations: GO: Gene ontology; KEGG: Kyoto Encyclopedia of Genes and Genomes; KIRC: Kidney renal clear cell carcinoma; MCODE: Molecular
complex detection; PPI: Protein-protein interaction.
CRL dysfunction. A link between malfunctioning Cullins represent a family of proteins that serve
mutations in CUL3 and Keap1 and papillary RCC has as a scaffold for E3 ubiquitin ligase and are linked to
been established in several investigations. In addition, tumor development. CUL1 has been identified as a key
12
the present study has identified a reduction in CUL3, player in several types of cancer, including lung cancer,
27
in conjunction with the substrate adaptors RhoBTB1 glioma, hepatocellular carcinoma, melanoma, gastric
29
28
30
and RhoBTB3, within the context of RCC. Recent cancer, and breast cancer. Studies have indicated that
31
32
25
developments in genomic technologies have facilitated its expression levels are associated with poor prognosis.
precise molecular characterization of cullin expression Previous research has shown that CUL2 serves primarily
patterns throughout the nephron, and transcriptome as a scaffolding protein for the E3 ubiquitin ligase during
26
analysis has demonstrated abundant expression of various viral infection. 33,34 Xu et al. observed significantly
30
cullins along the nephron segments. 9 elevated CUL2 expression in cervical cancer cells and
Volume 4 Issue 3 (2025) 8 doi: 10.36922/GPD025070014