Page 93 - IMO-2-2
P. 93
Innovative Medicines & Omics Femtomolar inhibition of pseudoeriocitrin
A B
Figure 5. Potential interactions between pseudoeriocitrin and rat carnitine palmitoyl transferase 2 (PDB ID: 2FW3). (A) The 3D representation of the
model. (B) The 2D representation of the model.
A B
Figure 6. Position of pseudoeriocitrin in carnitine palmitoyl transferase
2 (PDB ID: 2FW3). (A) In the representation of the enzyme surface,
the cavity in the orange circle was presumed to be the entry cavity of
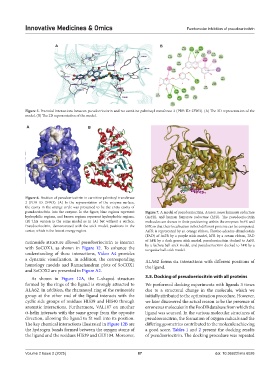
pseudoeriocitrin into the enzyme. In the figure, blue regions represent Figure 7. A model of pseudoeriocitrin, Ascaris suum fumarate reductase
hydrophilic regions, and brown regions represent hydrophobic regions. (AsFR), and human fumarate reductase (hFR). The pseudoeriocitrin
(B) This version is the same model as in (A) but without a surface. molecules are shown in their positioning within the enzymes AsFR and
Pseudoeriocitrin, demonstrated with the stick model, positions in the hFR so that their localization in both different proteins can be compared.
center, which is the lowest energy region. AsFR is represented by an orange ribbon, Flavine-adenine-dinucleotide
(FAD) of AsFR by a purple stick model, hFR by a cream ribbon, FAD
rutinoside structure allowed pseudoeriocitrin to interact of hFR by a dark green stick model, pseudoeriocitrin docked to AsFR
with SoCOX1, as shown in Figure 12. To enhance the by a fuchsia ball-stick model, and pseudoeriocitrin docked to hFR by a
turquoise ball-stick model.
understanding of these interactions, Video A4 provides
a dynamic visualization. In addition, the corresponding ALA62 forms six interactions with different positions of
homology models and Ramachandran plots of SoCOX1 the ligand.
and SoCOX2 are presented in Figure A2.
As shown in Figure 12A, the L-shaped structure 3.5. Docking of pseudoeriocitrin with all proteins
formed by the rings of the ligand is strongly attracted to We performed docking experiments with ligands 3 times
ALA62. In addition, the rhamnosyl ring of the rutinoside due to a structural change in the molecule, which we
group at the other end of the ligand interacts with the initially attributed to the optimization procedure. However,
cyclic side groups of residues HIS39 and HIS40 through we later discovered the actual reason to be the presence of
aromatic interactions. Furthermore, VAL107 on another erroneous molecules in the FooDB database from which the
α-helix interacts with the same group from the opposite ligand was sourced. In the various molecular structures of
direction, allowing the ligand to fit well into its position. pseudoeriocitrin, the formation of oxygen radicals and the
The key chemical interactions illustrated in Figure 12B are differing geometries contributed to the molecule achieving
the hydrogen bonds formed between the oxygen atoms of a good score. Tables 1 and 2 present the docking results
the ligand and the residues HIS39 and GLY104. Moreover, of pseudoeriocitrin. The docking procedure was repeated
Volume 2 Issue 2 (2025) 87 doi: 10.36922/imo.6026