Page 96 - IMO-2-2
P. 96
Innovative Medicines & Omics Femtomolar inhibition of pseudoeriocitrin
The selectivity index for pseudoeriocitrin was 4. Discussion
calculated to be 152,542, comparing the SoCOX1 enzyme 28
to the SoCOX2 enzyme based on their K values. The Recent in silico studies demonstrated that a new interaction
i
3D structures of both enzymes were predicted using can be formed within a morin molecule by calculating
homology modeling through the SwissModel web server. and comparing the enthalpy changes necessary for the
This in silico analysis indicates that pseudoeriocitrin is conformational change that enables an intramolecular
more effective at inhibiting the SoCOX1 enzyme than the bond. This new conformation allows morin to enhance
SoCOX2 enzyme. its radical scavenging activity while adopting a planar
structure. As a flavone, morin achieves a planar extension
Pseudoeriocitrin inhibits both AsFR and hFR homolog of its flavone core, leading to the formation of a ring, that
enzymes. In this case, the selectivity of pseudoeriocitrin is, parallel to this core. This condition mirrors the flavone
for nematode FR is lower than that for human FR, making structure examined in our study. The bonds that form
it less ideal as a ligand for FR inhibition. between the flavone core and the rutinoside group increase
Interestingly, pseudoeriocitrin was initially predicted the planar area made up of cyclic structures. Consequently,
to penetrate the blood-brain barrier, exhibit a high this arrangement enables the hydroxyl groups of the
absorption rate in the gastrointestinal tract, and bind to molecule to create a significant amount of hydrogen bonds
certain cytochrome enzymes that are not inhibited by with the target proteins, thereby enhancing its inhibitory
eriocitrin (SwissADME results for pseudoeriocitrin are effect.
shown in Figures 13 and A4). In the subsequent study on Another study demonstrated that modifications of
the same molecule, the SwissADME web server could not the isoleucyl-tRNA synthetase inhibitor mupirocin can
provide a prediction, as the molecular weight exceeded enhance its inhibitory effect. New inhibitors, developed
29
500 g/mol. The most significant finding in Figure 13A is based on modified mupirocin combined with an amino
that pseudoeriocitrin is identified as a multiradical. This acid side chain, demonstrated a much stronger inhibition
radical structure is associated with toxicity and binding of isoleucyl-tRNA synthetase, with dissociation constant
ability. However, pseudoeriocitrin is not present in the (K ) values of 10 – 12 fM, whereas the parent compound
d
ZincDatabase. Given its high free binding energy, the mupirocin inhibited the same enzyme with a K value of
d
interactions related to pseudoeriocitrin are intended to be 140 pM. Similarly, pseudoeriocitrin, a modified form
used in the discovery of new anthelmintics. of eriocitrin, demonstrates much higher inhibition
Table 2. Docking results of eriocitrin and pseudoeriocitrin (Ki values)
Molecule 1OJ0 2H4T 4YSX 6VAX SoCOX1 SoCOX2 6E7C Ev Tub CeGLUT1 2FW3
Eriocitrin - 663.49 nM 14.13 µM 370.31 pM 45.64 µM 1.34 mM - - 14.24 µM 75.23 nM
PE (in first docking) 2,270.00 µM 15.83 fM 71.76 pM 971.51 pM - 417.70 nM - - 479.82 nM 1.19 nM
PE (in second docking) - 55.00 fM 512.00 pM 29.86 fM 389.40 fM 59.40 nM - - 257.26 fM 3.45 fM
Notes: 1OJ0 represents Haemonchus contortus β-tubulin; 2FW3 represents rat carnitine o-palmitoyltransferase in complex with antidiabetic drug
ST1326; 2H4T represents rat carnitine o-palmitoyltransferase bound with dodecane; 4YSX represents Ascaris suum fumarate reductase; 6E7C
represents human β-tubulin; 6VAX represents human fumarate reductase.
Abbreviations: CeGLUT1: Caenorhabditis elegans Glucose transporter 1; EvTub: Enterobius vermicularis β-tubulin; PE: Pseudoeriocitrin;
SoCOX1: Syphacia obvelata cytochrome c oxidase 1; SoCOX2: Syphacia obvelata cytochrome c oxidase 2.
A B C
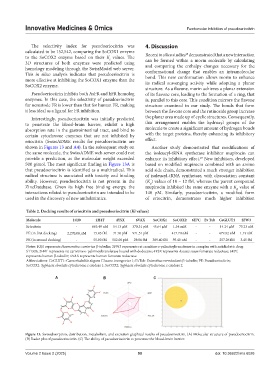
Figure 13. Swissabsorption, distribution, metabolism, and excretion graphical results of pseudoeriocitrin. (A) Molecular structure of pseudoeriocitrin.
(B) Radar plot of pseudoeriocitrin. (C) The ability of pseudoeriocitrin to penetrate the blood-brain barrier.
Volume 2 Issue 2 (2025) 90 doi: 10.36922/imo.6026