Page 102 - GPD-3-4
P. 102
Gene & Protein in Disease Pre-metastatic niche oral cancer: Insights
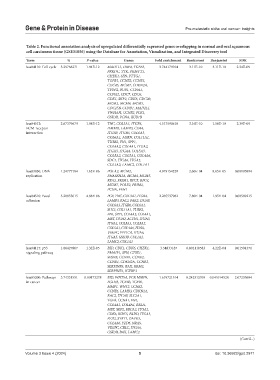
Table 2. Functional annotation analysis of upregulated differentially expressed genes overlapping in normal and oral squamous
cell carcinoma tissue (GSE31056) using the Database for Annotation, Visualization, and Integrated Discovery tool
Term % P‑value Genes Fold enrichment Bonferroni Benjamini FDR
hsa04110: Cell cycle 3.29768271 1.94E-12 MAD1L1, DBF4, TGFB3, 3.744179104 3.11E-10 3.11E-10 2.34E-09
PRKDC, TTK, PKMYT1,
CHEK1, SFN, PTTG1,
TGFB1, CCNE2, CCNE1,
CDC45, MCM7, CDKN2A,
TFDP2, BUB1, CCNA1,
CCNA2, CDC7, CDC6,
CDK1, SKP2, CDK6, CDC20,
MCM2, MCM4, MCM5,
CDC25B, CCNB1, MAD2L1,
YWHAH, CCNB2, PLK1,
GSK3B, PCNA, BUB1B
hsa04512: 2.67379679 1.98E-12 TNC, COL3A1, ITGB4, 4.517590618 3.16E-10 1.58E-10 2.39E-09
ECM-receptor HMMR, LAMB3, CD44,
interaction ITGAV, ITGB6, COL6A3,
COL6A1, AGRN, COL11A1,
THBS2, FN1, SPP1,
COL4A2, COL4A1, ITGA2,
ITGA3, ITGA4, COL5A3,
COL5A2, COL5A1, COL4A6,
SDC1, ITGA6, ITGA5,
COL1A2, LAMC2, COL1A1
hsa03030: DNA 1.24777184 1.62E-06 POLA2, MCM2, 4.919154229 2.60E-04 8.65E-05 0.00195894
replication RNASEH2A, MCM4, MCM5,
RPA3, PRIM1, RFC5, RFC4,
MCM7, POLE2, PRIM2,
PCNA, FEN1
hsa04510: Focal 3.20855615 4.88E-06 PGF, TNC, COL3A1, ITGB4, 2.265537982 7.80E-04 1.95E-04 0.00589215
adhesion LAMB3, RAC2, PAK2, ITGAV,
COL6A3, ITGB6, COL6A1,
SHC1, COL11A1, THBS2,
FN1, SPP1, COL4A2, COL4A1,
MET, ITGA2, ACTN1, ITGA3,
ITGA4, COL5A3, COL5A2,
COL5A1, COL4A6, FLNA,
VEGFC, PPP1CA, ITGA6,
ITGA5, GSK3B, COL1A2,
LAMC2, COL1A1
hsa04115: p53 1.60427807 1.32E-05 BID, CDK1, CDK6, CHEK1, 3.34833187 0.002110583 4.22E-04 0.01594371
signaling pathway PMAIP1, SFN, GTSE1,
SESN3, CCNB1, CCNE2,
CCNE1, CDKN2A, CCNB2,
SERPINB5, BAX, RRM2,
SERPINE1, IGFBP3
hsa05200: Pathways 3.74331551 0.00173278 BID, WNT5A, PGF, MMP9, 1.619721514 0.242313769 0.045194528 2.07235604
in cancer EGLN3, TGFB3, TGFB1,
MMP1, WNT2, CCNE2,
CCNE1, LAMB3, CDKN2A,
RAC2, ITGAV, SLC2A1,
TGFA, CCNA1, FN1,
COL4A2, COL4A1, RELA,
MET, SKP2, BRCA2, ITGA2,
CDK6, BIRC5, FADD, ITGA3,
FZD2, STAT1, DAPK3,
COL4A6, FZD6, NRAS,
VEGFC, CBLC, ITGA6,
GSK3B, BAX, LAMC2
(Cont’d...)
Volume 3 Issue 4 (2024) 5 doi: 10.36922/gpd.2971