Page 101 - GPD-3-4
P. 101
Gene & Protein in Disease Pre-metastatic niche oral cancer: Insights
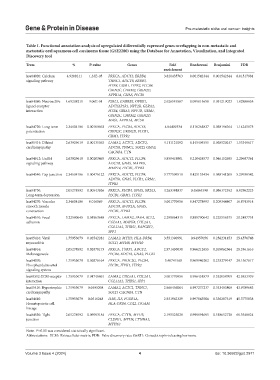
Table 1. Functional annotation analysis of upregulated differentially expressed genes overlapping in non‑metastatic and
metastatic oral squamous cell carcinoma tissue (GSE2280) using the Database for Annotation, Visualization, and Integrated
Discovery tool
Term % P‑value Genes Fold Bonferroni Benjamini FDR
enrichment
hsa04020: Calcium 4.9208211 1,33E-05 PRKCA, ADCY2, ERBB4, 3.820435763 0.001542344 0.001542344 0.01517981
signaling pathway TNNC1, ADCY8, ERBB3,
HTR4, GRM1, ITPR2, PLCB4,
GRIN2C, CHRM2, GRIN2D,
AVPR1A, GNAS, PLCB1
hsa04080: Neuroactive 4.69208211 9.06E-04 F2RL3, GABRB3, OPRK1, 2.626549587 0.099814658 0.051219023 1.02888658
ligand-receptor ADCYAP1R1, NPY2R, GLRA2,
interaction HTR4, GRIA3, NPY1R, GRM1,
GRIN2C, CHRM2, GRIN2D,
MAS1, AVPR1A, MC5R
hsa04720: Long-term 2.34604106 0.00100663 PRKCA, PLCB4, ADCY8, 4.94409334 0.110261827 0.038194164 1.14245075
potentiation GRIN2C, GRIN2D, PLCB1,
GRM1, ITPR2
hsa05414: Dilated 2.63929619 0.00135363 LAMA2, ACTC1, ADCY2, 4.111121092 0.145404555 0.038520247 1.53349617
cardiomyopathy ADCY8, TNNC1, SGCD, GNAS,
CACNB4, TTN
hsa04912: GnRH 2.63929619 0.00203465 PRKCA, ADCY2, PLCB4, 3.859419801 0.210424573 0.046152985 2.29687361
signaling pathway ADCY8, GNAS, MAPK8,
MMP14, PLCB1, ITPR2
hsa04540: Gap junction 2.34604106 0.00476122 PRKCA, ADCY2, PLCB4, 3.777509518 0.425135434 0.088141208 5.29938542
ADCY8, GNAS, PLCB1, GRM1,
ITPR2
hsa04730: 2.05278592 0.00543626 PRKCA, PLCB4, GNAS, GRIA3, 4.263384837 0.46864598 0.086372552 6.02962223
Long-term depression PLCB1, GRM1, ITPR2
hsa04270: Vascular 2.34604106 0.016069 PRKCA, ADCY2, PLCB4, 3.001770956 0.847278993 0.209344407 16.8743914
smooth muscle ADCY8, AVPR1A, GNAS,
contraction PLCB1, ITPR2
hsa04510: Focal 3.22580645 0.01963648 PRKCA, LAMA2, PAK4, BCL2, 2.299864315 0.899790642 0.225556375 20.2487751
adhesion COL6A1, MAPK8, COL2A1,
COL11A2, THBS1, RAPGEF1,
SPP1
hsa05416: Viral 1.75953079 0.02542281 LAMA2, MYH3, HLA‑DRB4, 3.551390991 0.94957056 0.258231453 25.4570748
myocarditis SGCD, MYH8, MYH10
hsa04916: 2.05278592 0.02878173 PRKCA, TYRP1, ADCY2, 2.971450038 0.966212655 0,265062564 28.3361616
Melanogenesis PLCB4, ADCY8, GNAS, PLCB1
hsa04070: 1.75953079 0.02976164 PRKCA, PIK3C2G, PLCB4, 3.40741568 0.969946202 0.253279147 29.1567617
Phosphatidylinositol PLCB1, ITPK1, ITPR2
signaling system
hsa04512: ECM-receptor 1.75953079 0.04746861 LAMA2, COL6A1, COL2A1, 3.001770956 0.996451879 0.352054709 42.5833939
interaction COL11A2, THBS1, SPP1
hsa05410: Hypertrophic 1.75953079 0.0495208 LAMA2, ACTC1, TNNC1, 2.966456004 0.997237237 0.343493808 43.9789683
cardiomyopathy SGCD, CACNB4, TTN
hsa04640: 1.75953079 0.0516248 IL9R, IL5, FCGR1A, 2.931962329 0.997863506 0.336287319 45.3775838
Hematopoietic cell HLA‑DRB4, CD22, ITGAM
lineage
hsa04530: Tight 2.05278592 0.09593184 PRKCA, CTTN, MYH3, 2.195325028 0.999991695 0.518652728 68.3548824
junction CLDN11, MYH8, CTNNA3,
MYH10
Note: P<0.05 was considered statistically significant.
Abbreviations: ECM: Extracellular matrix; FDR: False discovery rate; GnRH: Gonadotropin-releasing hormone.
Volume 3 Issue 4 (2024) 4 doi: 10.36922/gpd.2971