Page 104 - GPD-3-4
P. 104
Gene & Protein in Disease Pre-metastatic niche oral cancer: Insights
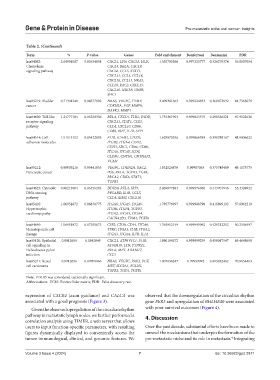
Table 2. (Continued)
Term % P‑value Genes Fold enrichment Bonferroni Benjamini FDR
hsa04062: 2.04991087 0.03634854 CXCL1, LYN, CXCL5, HCK, 1.555790566 0.997325777 0.326279374 36.0507054
Chemokine CXCL3, RELA, CXCL9,
signaling pathway CXCL6, CCL5, STAT1,
CXCL11, CCL4, CCL18,
CXCL10, CCL11, NRAS,
CCL20, RAC2, CXCL13,
CXCL16, GSK3B, GNB5,
SHC1
hsa05219: Bladder 0.71301248 0.04377802 NRAS, VEGFC, TYMP, 2.409381663 0.999224833 0.360872932 41.7563079
cancer CDKN2A, PGF, MMP9,
DAPK3, MMP1
hsa04620: Toll-like 1.24777184 0.05258336 RELA, CXCL9, TLR2, FADD, 1.753361903 0.999823575 0.398536221 47.9122636
receptor signaling CXCL11, STAT1, CCL5,
pathway CCL4, CXCL10, CD86,
CD80, IRF7, IL1B, SPP1
hsa04514: Cell 1.51515152 0.05422803 F11R, ICAM1, CD276, 1.629070556 0.999866393 0.390788167 48.9936622
adhesion molecules ITGB2, ITGA4, CDH2,
CDH3, SDC1, CD86, CD80,
ITGA6, ITGAV, ICOS,
CLDN1, CNTN1, CNTNAP2,
VCAN
hsa05212: 0.98039216 0.05441853 VEGFC, CDKN2A, RAC2, 1.932524876 0.99987063 0.375748489 49.1175775
Pancreatic cancer PGF, RELA, TGFB3, TGFA,
BRCA2, CDK6, STAT1,
TGFB1
hsa04623: Cytosolic 0.80213904 0.06456243 DDX58, RELA, IRF7, 2.069877883 0.999976966 0.413701946 55.3308922
DNA-sensing PYCARD, IL1B, CCL5,
pathway CCL4, AIM2, CXCL10
hsa05410: 1.06951872 0.06876775 ITGA6, ITGA5, ITGAV, 1.785776997 0.999988798 0.418899192 57.6962219
Hypertrophic ITGB6, ITGB4, TGFB3,
cardiomyopathy ITGA2, ITGA3, ITGA4,
CACNA2D3, TPM4, TGFB1
hsa04640: 1.06951872 0.07355873 CSF2, CD38, CD44, ITGA6, 1.765012149 0.999995092 0.426312252 60.2506497
Hematopoietic cell TFRC, ITGA5, IL1B, ITGA2,
lineage ITGA3, ITGA4, IL7R, IL1A
hsa05120: Epithelial 0.8912656 0.0842849 CXCL1, ATP6V1C1, F11R, 1.860184372 0.999999239 0.458017547 65.4648659
cell signaling in ADAM10, LYN, PTPRZ1,
Helicobacter pylori RELA, MET, ADAM17,
infection CCL5
hsa05211: Renal 0.8912656 0.09709066 NRAS, VEGFC, PAK2, PGF, 1.807036247 0.99999992 0.493832462 70.8654463
cell carcinoma MET, SLC2A1, EGLN3,
TGFB3, TGFA, TGFB1
Note: P<0.05 was considered statistically significant.
Abbreviations: ECM: Extracellular matrix; FDR: False discovery rate.
expression of CXCR4 (axon guidance) and CALCA was observed that the downregulation of the circadian rhythm
associated with a good prognosis (Figure 3). gene PER3 and upregulation of BHLHE40 were associated
Given the observed upregulation of the circadian rhythm with poor survival outcomes (Figure 4).
pathway in metastatic lymph nodes, we further performed a 4. Discussion
correlation analysis using TIMER, a web server that allows
users to input function-specific parameters, with resulting Over the past decade, substantial efforts have been made to
figures dynamically displayed to conveniently access the unravel the mechanisms that underpin the formation of the
tumor immunological, clinical, and genomic features. We pre-metastatic niche and its role in metastasis. Integrating
4
Volume 3 Issue 4 (2024) 7 doi: 10.36922/gpd.2971