Page 105 - GPD-3-4
P. 105
Gene & Protein in Disease Pre-metastatic niche oral cancer: Insights
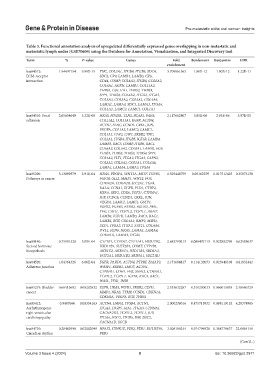
Table 3. Functional annotation analysis of upregulated differentially expressed genes overlapping in non‑metastatic and
metastatic lymph nodes (GSE70604) using the Database for Annotation, Visualization, and Integrated Discovery tool
Term % P‑value Genes Fold Bonferroni Benjamini FDR
enrichment
hsa04512: 1.94497154 9.89E-15 TNC, COL3A1, ITGB4, ITGB5, SDC4, 3.709961563 1.80E-12 1.80E-12 1.22E-11
ECM-receptor SDC2, GP9, LAMB4, LAMB3, GP6,
interaction CD44, COMP, COL6A3, ITGB6, COL6A2,
COL6A1, AGRN, LAMB1, COL11A2,
THBS1, COL11A1, THBS2, THBS3,
SPP1, THBS4, COL4A2, ITGA2, ITGA3,
COL5A3, COL5A2, COL5A1, COL4A6,
LAMA1, LAMA4, SDC1, LAMA3, ITGA6,
COL1A2, LAMC2, LAMC1, COL1A1
hsa04510: Focal 2.65654649 3.22E-08 HRAS, PDGFA, TLN2, BCAR1, PAK6, 2.117662807 5.85E-06 2.93E-06 3.97E-05
adhesion COL11A2, COL11A1, EGFR, ACTN4,
ACTN1, FLNC, CCND1, CRKL, JUN,
VEGFA, COL1A2, LAMC2, LAMC1,
COL1A1, CAV2, CAV1, ERBB2, TNC,
COL3A1, ITGB4, ITGB5, IGF1R, LAMB4,
LAMB3, RAC3, COMP, ITGB6, RAC1,
COL6A3, COL6A2, COL6A1, LAMB1, EGF,
THBS1, THBS2, THBS3, THBS4, SPP1,
COL4A2, FLT1, ITGA2, ITGA3, CAPN2,
COL5A3, COL5A2, COL5A1, COL4A6,
LAMA1, LAMA4, LAMA3, ITGA6
hsa05200: 3.13092979 2.91E-04 HRAS, PDGFA, WNT3A, MITF, TGFB3, 1.529448759 0.05162259 0.017512425 0.35873138
Pathways in cancer FGF10, GLI2, MMP1, WNT2, FOS,
CDKN2A, CDKN2B, SLC2A1, TGFA,
RALA, CCNA1, EGFR, PLD1, CTBP2,
RXRA, SKP2, CDK6, FGF21, CTNNA1,
JUP, CCDC6, CCND1, CRKL, JUN,
VEGFA, LAMC2, LAMC1, GSTP1,
FGFR2, FGFR3, ERBB2, EGLN3, PML,
TFG, CDH1, TCF7L2, TCF7L1, ARNT,
LAMB4, IGF1R, LAMB3, RAC3, RAC1,
LAMB1, EGF, COL4A2, BMP2, MSH3,
FZD1, ITGA2, ITGA3, STAT1, COL4A6,
DVL1, FZD6, NRAS, LAMA1, LAMA4,
CDKN1A, LAMA3, ITGA6
hsa00140: 0.75901328 5.09E-04 CYP1B1, CYP3A7, CYP11A1, HSD17B2, 2.643790213 0.088437113 0.022882798 0.62585637
Steroid hormone HSD17B1, SULT2B1, COMT, CYP7B1,
biosynthesis AKR1C2, AKR1C4, HSD11B1, SRD5A3,
UGT2A1, HSD11B2, SRD5A1, SULT1E1
hsa04520: 1.04364326 6.66E-04 EGFR, PARD3, ACTN4, PTPRF, BAIAP2, 2.171684817 0.114132053 0.023946108 0.81832442
Adherens junction WASF1, ERBB2, LMO7, ACTN1,
CTNND1, CDH1, FER, SNAI2, CTNNA1,
TCF7L2, TCF7L1, IGF1R, RAC3, RAC1,
WASL, YES1, INSR
hsa05219: Bladder 0.66413662 0.00203832 EGFR, HRAS, FGFR3, ERBB2, CDH1, 2.533632287 0.310200033 0.060015855 2.48646729
cancer MMP1, NRAS, TYMP, CCND1, CDKN1A,
CDKN2A, VEGFA, EGF, THBS1
hsa05412: 0.9487666 0.00354163 ACTN4, LMNA, ITGB4, ACTN1, 2.000236016 0.475715932 0.088119152 4.28374905
Arrhythmogenic ITGA2, ITGB5, GJA1, ITGA3, CTNNA1,
right ventricular CACNA2D2, TCF7L2, TCF7L1, JUP,
cardiomyopathy ITGA6, SGCG, ITGB6, DSP, DSC2,
CACNA1D, SGCB
hsa04710: 0.28462998 0.02002098 NPAS2, CSNK1E, PER2, PER1, BHLHE40, 3.508106244 0.974796926 0.368778637 22.0864146
Circadian rhythm PER3
(Cont’d...)
Volume 3 Issue 4 (2024) 8 doi: 10.36922/gpd.2971