Page 134 - GPD-3-4
P. 134
Gene & Protein in Disease A pyroptosis-related gene signature in myeloma
A
B
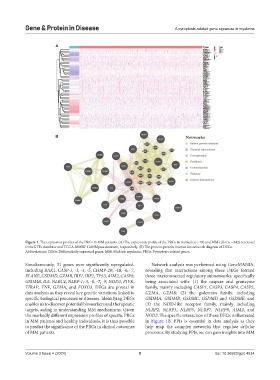
Figure 1. The expression profiles of the PRGs in MM patients. (A) The expression profile of the PRGs in normal (n = 70) and MM cells (n = 842) retrieved
from GTEx database and TCGA-MMRF CoMMpass database, respectively. (B) The protein-protein interaction network diagram of DEGs.
Abbreviations: DEGs: Differentially expressed genes; MM: Multiple myeloma; PRGs: Pyroptosis-related genes.
Simultaneously, 31 genes were significantly upregulated, Network analysis was performed using GeneMANIA,
including BAK1, CASP-1, -3, -4, -5, CHMP-2B, -4B, -6,- 7, revealing that interactions among these DEGs formed
ELANE, GSDMD, GZMB, IRF1, IRF2, TP53, AIM2, CASP8, three interconnected regulatory subnetworks, specifically
GSDMB, IL6, NLRC4, NLRP-1,-3, -6, -7, -9, NOD2, PJVK, being associated with: (1) the caspase and granzyme
TIRAP, TNF, GZMA, and FOXO3. DEGs are pivotal in family, mainly including CASP1, CASP3, CASP4, CASP5,
data analysis as they reveal key genetic variations linked to GZMA, GZMB; (2) the gsdermin family, including
specific biological processes or diseases. Identifying DEGs GSDMA, GSDMB, GSDMC, GSDMD and GSDME; and
enables us to discover potential biomarkers and therapeutic (3) the NOD-like receptor family, mainly including
targets, aiding in understanding MM mechanisms. Given NLRP2, NLRP3, NLRP5, NLRP7, NLRP9, AIM2, and
the markedly different expression profiles of specific PRGs NOD2. The specific interaction of these DEGs is illustrated
in MM patients and healthy individuals, it is thus possible in Figure 1B. PPIs is essential in data analysis as they
to predict the significance of the PRGs in clinical outcomes help map the complex networks that regulate cellular
of MM patients. processes. By studying PPIs, we can gain insights into MM
Volume 3 Issue 4 (2024) 5 doi: 10.36922/gpd.4534