Page 36 - GPD-4-3
P. 36
Gene & Protein in Disease TNFA polymorphism and risk of endometriosis
Table 6. Subgroup analysis of the ‑238 G>A variant based on ethnicity
Model Ethnicity Number of Test of association Test of heterogeneity Publication bias
studies OR 95% CI P‑value Adjusted Model P‑value I 2 P‑value
P value (Egger’s test)
Allele Asian 4 0.98 0.6893 – 1.3960 0.914958 1 Fixed 0.1972 0.3582 0.9894
Caucasian 1 0.85 0.3009 – 2.4050 0.760328 1 Fixed NA NA NA
Recessive Asian 3 1.36 0.5039 – 3.6581 0.545358 1 Fixed 0.4822 0 0.0627
Caucasian 0 NA
Dominant Asian 4 0.88 0.4628 – 1.6656 0.690448 1 Random 0.086 0.5451 0.6024
Caucasian 1 0.84 0.2905 – 2.4498 0.754432 1 Fixed NA NA NA
OD Asian 4 0.80 0.3802 – 1.6683 0.546315 1 Random 0.0503 0.6154 0.4711
Caucasian 1 0.84 0.2905 – 2.4498 0.754432 1 Fixed NA NA NA
HR vs. HW Asian 3 1.10 0.3945 – 3.0785 0.852951 1 Fixed 0.3748 0 0.0454
Caucasian 0 NA
HR vs. HT Asian 3 2.04 0.7079 – 5.8749 0.186825 1 Fixed 0.3637 0.0114 0.2229
Caucasian 0 NA
HT vs. HW Asian 4 0.79 0.3684 – 1.6778 0.534033 1 Random 0.0465 0.624 0.4428
Caucasian 1 0.84 0.2905 – 2.4498 0.754432 1 Fixed NA NA NA
Abbreviations: CI: Confidence interval; HR: Homologous recombination; HT: Heterozygote; HW: Homozygous wild; NA: Not available;
OD: Overdominant; OR: Odds ratio; vs.: Versus.
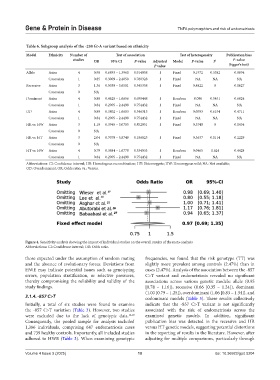
Figure 4. Sensitivity analysis showing the impact of individual studies on the overall results of the meta-analysis
Abbreviations: CI: Confidence interval; OR: Odds ratio.
those expected under the assumption of random mating frequencies, we found that the risk genotype (TT) was
and the absence of evolutionary forces. Deviations from slightly more prevalent among controls (2.47%) than in
HWE may indicate potential issues such as genotyping cases (2.47%). Analysis of the association between the -857
errors, population stratification, or selective pressures, C>T variant and endometriosis revealed no significant
thereby compromising the reliability and validity of the associations across various genetic models: allele (0.95
study findings. [0.78 – 1.16]), recessive (0.66 [0.35 – 1.24]), dominant
(1.00 [0.79 – 1.25]), overdominant (1.06 [0.83 – 1.34]), and
3.1.4. -857 C>T codominant models (Table 5). These results collectively
Initially, a total of six studies were found to examine indicate that the -857 C>T variant is not significantly
the -857 C>T variation (Table 3). However, two studies associated with the risk of endometriosis across the
were excluded due to the lack of genotypic data. 12,18 examined genetic models. In addition, significant
Consequently, the pooled sample for analysis included publication bias was detected in the recessive and HR
1,386 individuals, comprising 647 endometriosis cases versus HT genetic models, suggesting potential distortions
and 739 healthy controls. Importantly, all included studies in the reporting of results in the literature. However, after
adhered to HWE (Table 3). When examining genotypic adjusting for multiple comparisons, particularly through
Volume 4 Issue 3 (2025) 10 doi: 10.36922/gpd.5204