Page 44 - GTM-1-1
P. 44
Global Translational Medicine Bioinformatics analysis of dilated cardiomyopathy
A B
C
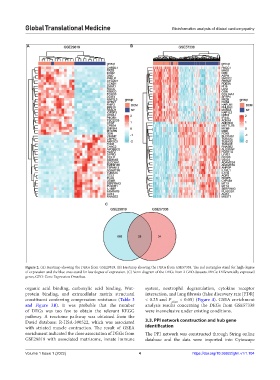
Figure 2. (A) Heatmap showing the DEGs from GSE29819. (B) Heatmap showing the DEGs from GSE57338. The red rectangles stand for high degree
of expression and the blue ones stand for low degree of expression. (C) Venn diagram of the DEGs from 2 GEO datasets. DEGs: Differentially expressed
genes, GEO: Gene Expression Omnibus.
organic acid binding, carboxylic acid binding, Wnt- system, neutrophil degranulation, cytokine receptor
protein binding, and extracellular matrix structural interaction, and lung fibrosis (false discovery rate [FDR]
constituent conferring compression resistance (Table 3 < 0.25 and P adjust < 0.05) (Figure 4). GSEA enrichment
and Figure 3B). It was probable that the number analysis results concerning the DEGs from GSE57338
of DEGs was too few to obtain the relevant KEGG were inconclusive under existing conditions.
pathway. A reactome pathway was obtained from the
David database: R-HSA-390522, which was associated 3.3. PPI network construction and hub gene
with striated muscle contraction. The result of GSEA identification
enrichment indicated the close association of DEGs from The PPI network was constructed through String online
GSE29819 with associated matrisome, innate immune database and the data were imported into Cytoscape
Volume 1 Issue 1 (2022) 4 https://doi.org/10.36922/gtm.v1i1.104