Page 110 - IMO-2-2
P. 110
Innovative Medicines & Omics Modeling Aurora-B kinase inhibitors
Table 2. Statistical parameters of the best AADRR model
Hypothesis ID PLS Factor SD R 2 F P Stability RMSE Q 2 Pearson R
AADRR.94 1 0.3668 0.8284 183.5 3.999 e-16 0.9384 0.357 0.804 0.8989
2 0.2337 0.9322 254.3 2.389 e-22 0.8595 0.2609 0.8953 0.9463
3 0.1547 0.9711 403 9.626 e-28 0.7662 0.245 0.9077 0.9529
Notes: PLS: Partial least square; SD: Standard deviation of the regression; R : Coefficient of determination; F: Variance ratio; P: Significance level of
2
the variance ratio; RMSE: Root mean square error; Q : Cross-validated correlation coefficient; Pearson R: correlation between predicted and observed
2
activity for test set.
A B
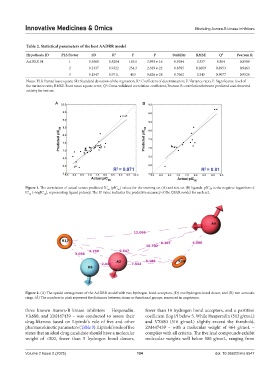
Figure 1. The correlation of actual versus predicted IC (pIC ) values for the training set (A) and test set (B) ligands. pIC₅₀ is the negative logarithm of
50
50
IC (−logIC ), representing ligand potency. The R value indicates the predictive accuracy of the QSAR model for each set.
2
50 50
Figure 2. (A) The spatial arrangement of the AADRR model with two hydrogen-bond acceptors, (D) one hydrogen-bond donor, and (R) two aromatic
rings. (Å) The numbers in pink represent the distances between atoms or functional groups, measured in angstroms.
three known Aurora-B kinase inhibitors – Hesperadin, fewer than 10 hydrogen bond acceptors, and a partition
VX680, and ZM447439 – was conducted to assess their coefficient (log P) below 5. While Hesperadin (513 g/moL)
drug-likeness based on Lipinski’s rule of five and other and VX680 (516 g/moL) slightly exceed the threshold,
pharmacokinetic parameters (Table 3). Lipinski’s rule of five ZM447439 – with a molecular weight of 464 g/moL –
states that an ideal drug candidate should have a molecular complies with all criteria. The five lead compounds exhibit
weight of <500, fewer than 5 hydrogen bond donors, molecular weights well below 500 g/moL, ranging from
Volume 2 Issue 2 (2025) 104 doi: 10.36922/imo.6547