Page 115 - IMO-2-2
P. 115
Innovative Medicines & Omics Modeling Aurora-B kinase inhibitors
Table 4. The key residues of protein‑ligand interactions identified for the lead compounds in molecular docking study and MD
simulations
Lead compound IUPAC name Molecular MD simulations
(NCI ID) docking
1 (695163) (E)-1-(1H-benzo[d] imidazol-2-yl)-3-(4-methoxyphenyl) ALA 157 LYS106, ALA157, ASN205, GLU204
prop-2-en-1-one
2 (327359) (E)-2-(((2-((2-hydroxyethyl) thio) benzo[d] thiazol-6-yl) LYS106, PHE219 GLU161, LYS106
imino) methyl) phenol
3 (721045) (E)-9-(2-(4-chlorophenyl)-2-(hydroxyimino) ethoxy) ALA157 ALA157, GLU161, LEU83
chromeno[3,4-b] chromene-6,12-dione
4 (711797) (E)-3-(4-(benzo[d][1,3]dioxol-5-yl)-2-oxobut-3-en-1-yl) ALA157, PHE219, ALA157, PHE88, GLU204, LYS106
pyrido[2,3-b] pyrazin-2 (1H)-one LYS106.
5 (104546) 4-((E)-((E)-(8-hydroxyquinolin-2 (1H)-ylidene) hydrazono) LYS106, PHE219, ALA157, GLU155, GLU204, GLY84, LYS202
methyl) quinolin-8-ol PHE88.
Abbreviation: MD: Molecular dynamics.
A B
C D
E
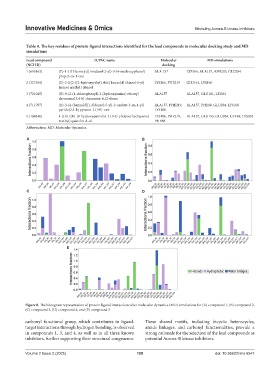
Figure 8. The histogram representation of protein-ligand interactions after molecular dynamics (MD) simulations for (A) compound 1, (B) compound 2,
(C) compound 3, (D) compound 4, and (E) compound 5
carbonyl functional group, which contributes to ligand- These shared motifs, including bicyclic heterocycles,
target interactions through hydrogen bonding, is observed amide linkages, and carbonyl functionalities, provide a
in compounds 1, 3, and 4, as well as in all three known strong rationale for the selection of the lead compounds as
inhibitors, further supporting their structural congruence. potential Aurora-B kinase inhibitors.
Volume 2 Issue 2 (2025) 109 doi: 10.36922/imo.6547