Page 115 - MI-1-1
P. 115
Microbes & Immunity Comprehensive genomic surveillance analysis of SARS-CoV-2
in submission lag times decreased over time for both We further analyzed the fold increase in reproductive
continents and VOCs/VOIs. The shortest submission lag number against the date of lineage emergence over time
times for continents and VOCs/VOIs were Europe and (as of August 15, 2022) based on publicly released data
Omicron, respectively. provided by the Broad Institute of MIT and Harvard
(http://www.broadinstitute.org/) in the pyro-cov data
The cumulative number of cases, cumulative number of repository (https://github.com/broadinstitute/pyro-cov).
7
deaths, and mortality rates of SARS-CoV-2 over time (as of As shown in Figure 5, the overall mean fold increase in
September 30, 2022) were analyzed using publicly available reproductive number for SARS-CoV-2 was 3.26 (median:
data from the Center for Systems Science and Engineering 1.16; 95% CI: 0.84 – 17.0). The mean fold increase in
(CSSE) at Johns Hopkins University in the COVID-19 reproductive number for Alpha, Beta, Gamma, Delta,
Data repository (https://github.com/CSSEGISandData/ Omicron, Epsilon, Iota, and Others was 1.65 (median:
COVID-19). As shown in Figure 4, the overall trends 1.64; 95% CI: 1.56 – 1.70), 1.57 (median: 1.56; 95% CI:
6
in the cumulative number of cases and deaths exhibited 1.53 – 1.62), 1.91 (median: 1.88; 95% CI: 1.78 – 2.04), 2.63
similar patterns over time. The epidemic commenced with (median: 2.62; 95% CI: 2.26 – 2.99), 13.22 (median: 12.06;
a rapid surge in cases, followed by gradual deceleration, 95% CI: 6.02 – 20.59), 1.38 (median: 1.35; 95% CI: 1.34 –
and ultimately reaching a stable phase by September 30, 1.44), 1.54 (95% CI: 1.32 – 1.80), and 1.15 (median: 1.10;
2022, the final date included in our study. The overall mean 95% CI: 0.89 – 1.50), respectively. The mean fold increase in
mortality rate of SARS-CoV-2 is 2.62% (median: 2.17%; reproductive number for non-Omicron variants was 1.40
95% CI: 1.06 – 6.81%). Notably, the highest mortality rate (median: 1.12; 95% CI: 0.97 – 2.28). These findings suggest
(7.73%) was recorded on April 29, 2020, preceding the that VOC Omicron exhibits distinct transmissibility
emergence of VOCs/VOIs, while the lowest mortality rate characteristics compared to previous SARS-CoV-2 strains.
(1.06%) was recorded on September 30, 2022, subsequent This increased transmissibility is likely attributable to its
to the emergence of Omicron and its highly contagious numerous mutations, which have enabled it to spread
subvariants (Videoclip S1). more efficiently, 8-12 as evidenced by its highest recorded
fold increase in reproductive number (24.15).
A Our study also acknowledges several limitations. First,
it is important to note that not all COVID-19 cases are
reported, sampled, and sequenced. Second, the cases that
are reported, sampled, and sequenced may not be entirely
representative of the broader situation within each country,
particularly in LMICs. Third, it is crucial to recognize
B that not all sequenced cases have high-quality datasets or
available metadata.
The study, conducted through extensive metadata
analysis, has revealed significant geographic variability in the
distribution of VOCs/VOIs across different continents, with
a notable emphasis on VOIs Epsilon and Iota. In addition,
considerable diversity in sequencing turnaround times has
C been observed across continents, with Europe displaying the
shortest times among all continents and the Omicron variant
demonstrating the swiftest genomic surveillance among
VOCs/VOIs. Despite the lower mortality rate associated
with the Omicron variant, its heightened transmissibility
in comparison to earlier SARS-CoV-2 strains underscores
the continued need for comprehensive strategies and
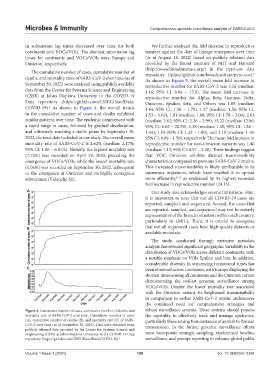
Figure 4. Cumulative number of cases, cumulative number of deaths, and robust surveillance systems. These systems should possess
mortality rate of SARS-CoV-2 over time. Cumulative number of cases the capability to effectively track and manage epidemics,
(A), cumulative number of deaths (B), and mortality rate (C) of SARS- particularly those arising from instances of animal-to-human
CoV-2 over time (as of September 30, 2022). Data were obtained from transmission. In the future, genomic surveillance efforts
publicly released data provided by the Center for Systems Science and
Engineering (CSSE) at Johns Hopkins University in the COVID-19 Data must incorporate strategic sampling, randomized baseline
repository (https://github.com/CSSEGISandData/COVID-19). 6 surveillance, and prompt reporting to enhance global public
Volume 1 Issue 1 (2024) 109 doi: 10.36922/mi.2294