Page 65 - MI-2-2
P. 65
Microbes & Immunity Phylogenetic analysis of HPV16 L1 in Asia
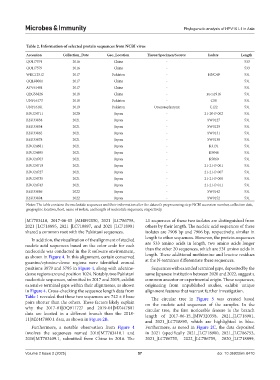
Table 2. Information of selected protein sequences from NCBI virus
Accession Collection_Date Geo_Location Tissue/Specimen/Source Isolate Length
QOI17574 2016 China - 533
QOI17579 2016 China - 533
WKC12512 2017 Pakistan - HNC49 531
QQL88061 2017 China - 531
AYV61481 2017 China - 531
QEG53826 2018 China - xuca1916 531
UNF16173 2018 Pakistan - C50 531
UNF16181 2019 Pakistan Oronasopharynx C122 531
BDO24711 2020 Japan - 21-20-P-002 531
BEU33838 2021 Japan - SW0127 531
BEU33854 2021 Japan - SW0129 531
BEU33862 2021 Japan - SW0131 531
BEU33878 2021 Japan - SW0138 531
BDO24681 2021 Japan - K3131 531
BDO24695 2021 Japan - K5048 531
BDO24703 2021 Japan - K5060 531
BDO24719 2021 Japan - 21-21-P-001 531
BDO24727 2021 Japan - 21-21-P-007 531
BDO24735 2021 Japan - 21-21-P-008 531
BDO24743 2021 Japan - 21-21-P-011 531
BEU33886 2022 Japan - SW0142 531
BEU33894 2022 Japan - SW0152 531
Notes: The table contains the nucleotide sequences and their information after the dataset’s preprocessing step: NCBI accession number, collection date,
geographic location, host, name of isolate, and length of nucleotide sequences, respectively.
|MT783410, 2017-06-05 |MH892050, 2021 |LC786758, L1 sequences of these two isolates are distinguished from
2021 |LC718895, 2021 |LC718897, and 2021 |LC718901 others by their length. The nucleic acid sequences of these
shared a common root with the Pakistani sequences. isolates are 7908 bp and 7906 bp, respectively, similar in
In addition, the visualization of the alignment of studied length to other sequences. However, the protein sequences
nucleic acid sequences based on the color code for each are 533 amino acids in length, two amino acids longer
nucleotide was conducted in the R software environment, than the other 20 sequences, which are 531 amino acids in
as shown in Figure 4. In this alignment, certain conserved length. These additional methionine and leucine residues
guanine/cytosine-dense regions were identified around at the N-terminus differentiate these sequences.
positions 3979 and 5795 in Figure 4, along with adenine- Sequences with extended terminal gaps, deposited by the
dense regions around position 1024. Notably, two Pakistani same Japanese institution between 2020 and 2022, suggest a
nucleotide sequences, submitted in 2017 and 2019, exhibit common ancestor or experimental origin. These sequences,
extensive terminal gaps within their alignments, as shown originating from unpublished studies, exhibit unique
in Figure 4. Cross-checking the sequence length data from alignment features that warrant further investigation.
Table 1 revealed that these two sequences are 742 ± 8 base The circular tree in Figure 5 was created based
pairs shorter than the others. These factors likely explain on the nucleic acid sequences of the samples. In the
why the 2017-01|OQ911727 and 2019-01|MZ447801 circular tree, the first noticeable feature is the branch
data are located in a different branch than the 2018- length of 2017-06-15_|MW320358, 2021_|LC718901,
11|MZ447800.1 data, as shown in Figure 2B. and 2021_|LC718895, which are highlighted in blue.
Furthermore, a notable observation from Figure 4 Furthermore, as noted in Figure 2C, the data deposited
involves the sequences named 2016|MT783410.1 and in 2021 (specifically 2021_|LC718900, 2021_|LC786753,
2016|MT783409.1, submitted from China in 2016. The 2021_|LC786755, 2022_|LC786759, 2020_|LC718899,
Volume 2 Issue 2 (2025) 57 doi: 10.36922/mi.8410